Our recent research paper published in Nature Partner Journals describes the development of an AI model trained on large-scale MRI scans and radiology reports.1 This research explores how a single model may be adapted to support tasks such as detecting patterns associated with disease, mapping anatomy, identifying potential abnormalities, and linking scans with reports without requiring full system retraining for each use case.
Decipher-MR is a 3D MRI-specific vision-language foundation model pretrained on more than 200,000 MRI series from over 22,000 studies. Instead of building a separate large model for each use case, this research approach keeps the core MRI model fixed and evaluates the addition of smaller task-specific components for classification, retrieval, segmentation, localization, and text-guided search.
A key area of investigation in this work is the development of a shared MRI model that may transfer across body regions, sequences, protocols, and task types. This approach is being explored as a way to support more modular and potentially more efficient MRI AI development, with the possibility of extending to additional research applications.
The challenge
MRI is one of the most widely used imaging modalities in clinical diagnosis and research, but it is also one of the most complex to analyze. Unlike more standardized image types, MRI varies across scanner hardware, pulse sequences, protocols, anatomical regions, and clinical contexts. MRI is also inherently three-dimensional, requiring models to learn from full scan volumes rather than two-dimensional images.
As a result, many MRI AI approaches continue to be developed for narrow, task-specific applications and often rely on labeled datasets that can be resource-intensive to develop at scale.
Foundation models have contributed to advances in language and computer vision, and similar approaches are being explored in medical imaging. However, MRI remains relatively underrepresented in many multimodal medical models, where it may constitute only a small portion of pretraining data. Other MRI-focused approaches are often limited to specific anatomical regions, such as brain-only training, which may reduce generalizability.
This highlights an opportunity for research into MRI-specific models trained across broader datasets, including multiple body regions, imaging sequences, and clinical contexts.
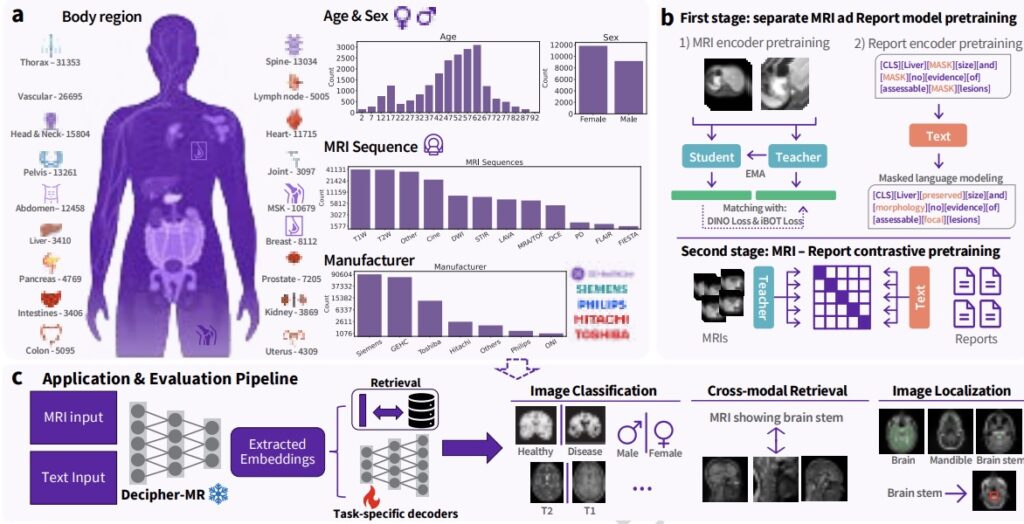
A reusable 3D foundation model built specifically for MRI
To explore these challenges, Decipher-MR was pretrained on a large MRI-only dataset comprising 22,594 studies and 203,233 MRI series, with radiology reports available for 20,658 studies. The dataset spans ages 0 to 90, includes a range of body regions and MRI protocols, and incorporates scans from multiple vendors.
Rather than treating MRI as a subset within a broader imaging dataset, this research focuses on MRI-specific characteristics, including volumetric structure, protocol variability, and use across diverse clinical workflows.
The model architecture is designed for reuse. Instead of retraining large encoders for each downstream task, the research evaluates a configuration where the core model remains fixed and smaller task-specific components are added for classification, retrieval, segmentation, localization, and grounding. This approach investigates whether a single pretrained MRI backbone may support multiple task types across the body.
Methodology
Decipher-MR is trained using both MRI scans and associated radiology reports. This enables the model to learn relationships between imaging features and the clinical language used to describe anatomy, findings, and disease.
This combined image-and-text training is intended to support linking scans with report text, distinguishing patterns in anatomy and abnormalities, and responding to text-based queries about imaging content. Additional methodological details are available in the published research.
Benchmark performance
The model was evaluated in a frozen-encoder setting across classification, regression, retrieval, segmentation, localization, and visual grounding tasks. The goal of this evaluation was to assess whether a shared pretrained MRI backbone could be reused across different task categories with lightweight downstream components.
Across these evaluations, Decipher-MR demonstrated consistent performance across task types and showed improvements relative to selected comparison models under the reported conditions. Performance was also evaluated in lower-data settings.
For example, in full-report retrieval tasks, the model identified the paired scan within the top 10 results in approximately 26% of cases, compared with 5.1% for MedImageInsight under the same reported conditions.
We also assessed performance across male and female cohorts. While variation was observed, the model showed relatively consistent performance across groups compared to the selected benchmarks.
Classification and regression
Across the reported classification tasks, Decipher-MR demonstrated improvements relative to DINOv2, BiomedCLIP, and, in most cases, MedImageInsight within the evaluated setup. One example included an average improvement of 2.9% in disease classification compared to MedImageInsight.
It is important to note that performance gains were not uniform across all datasets and tasks. In some cases, results were similar to or favored baseline models. Preprocessing methods also influenced results, indicating potential areas for further optimization.
The importance of text supervision and data diversity
Ablation studies suggest that training with both MRI scans and radiology reports contributed to observed performance across several tasks. For example, a 4.2% improvement in body region detection was observed in lower-data settings.
Pretraining on a diverse MRI dataset also demonstrated improved performance compared to models trained on narrower datasets, supporting the importance of both data diversity and report-based supervision.
Cross-modal retrieval
Because the model is trained on both imaging and text data, it can link MRI scans with associated report content. In an out-of-domain body-region retrieval task, top-3 success reached 91.4% using full reports, compared with 81.1% for MedImageInsight under the same reported conditions. Performance was stronger for anatomy-focused queries than for pathology-focused queries, highlighting an area for continued research.
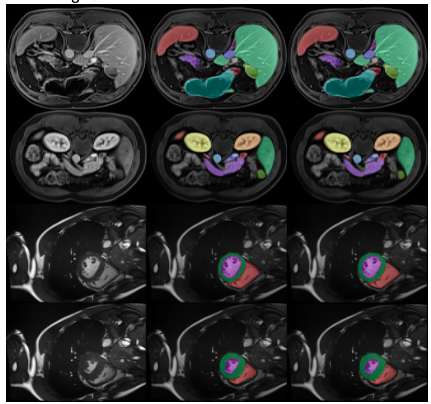
Segmentation
For 3D anatomical segmentation, Decipher-MR paired with a ResNet-based decoder demonstrated performance exceeding SAMMed3D under the same frozen-encoder conditions and produced results comparable to nnU-Net in the reported experiments. One reported result included achieving a Dice score of 0.82 on ACDC within one epoch, suggesting potential for efficient transfer learning under the evaluated conditions.
Localization and visual grounding
The model was also evaluated for anomaly localization and text-guided visual grounding tasks, such as identifying regions of interest or anatomical structures. Replacing visual encoders in localization frameworks with a frozen Decipher-MR encoder resulted in a reported 14% performance improvement under the tested conditions.
Conclusion
Decipher-MR represents ongoing research into MRI foundation modeling, combining large-scale 3D MRI pretraining, report-guided text supervision, and a modular adaptation approach. The results suggest that a shared MRI-focused backbone may be capable of supporting multiple task categories, body regions, and imaging contexts within a research setting.
- Concept only. This material describes early-stage research and is provided for informational and exploratory purposes. The technology described is under investigation, may change, and may never become a product. Not for sale. Not cleared or approved by the U.S. FDA or any other regulatory authority for commercial availability. Any results described are preliminary and subject to further validation. ↩︎