An MRI method may work well in a controlled study, for example by identifying the scan type, checking whether an image was acquired correctly, or correcting a distorted image. An important question is whether it remains reliable in real clinical settings, where patients differ, scan settings vary, and image quality is not always ideal.
That was the broader focus of the research work we presented at ISMRM 2026. Across these studies, we explored reliability at several points in the MRI workflow, from understanding what kind of image is being analyzed, to checking whether a scan was prescribed correctly, to supporting setup of a cardiac exam, aligning images across scans, and correcting distortion in fast imaging.
Taken together, our research1 reflects a shift away from narrow model performance and toward methods that are intended to be more dependable in everyday imaging conditions.
Preparing MRI images for more reliable AI
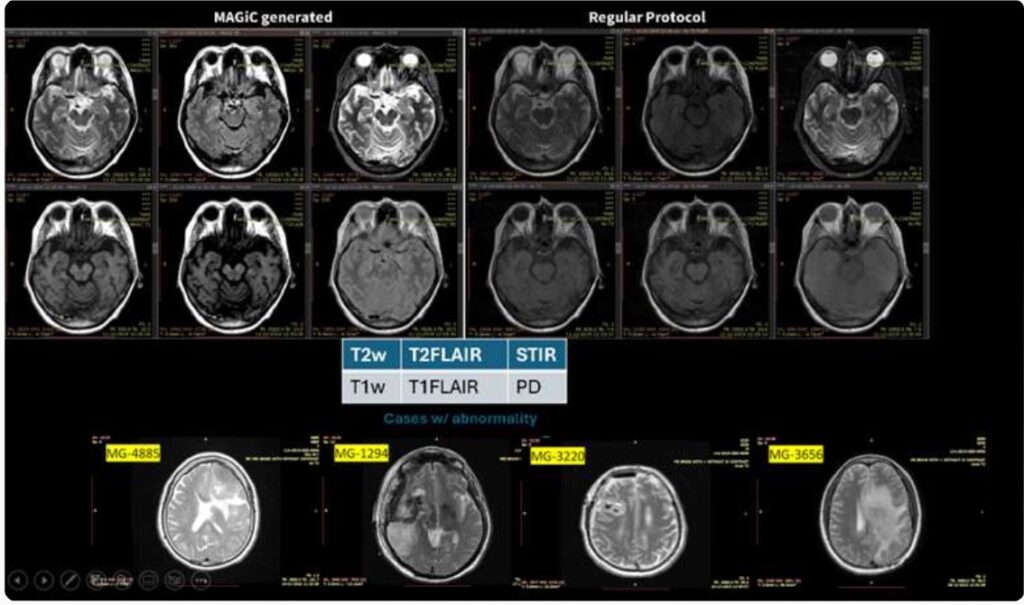
Foundation models are large models trained to produce reusable image features. In simple terms, they turn an image into a compact summary that other tools can use for different tasks. This approach is of interest in MRI, where many tasks depend on the same anatomy, but it also raises an important question: what is an appropriate way to prepare an MRI scan before those features are extracted?
We studied that question using a 3D MR foundation model on two basic but important tasks. The first was contrast identification, meaning recognizing what type of scan it is, such as T1, T2, or FLAIR. The second was orientation identification, meaning recognizing whether the image is viewed from above, from the side, or from the front. These checks can influence downstream tools that depend on them being correct.
To evaluate this, we compared several ways of preparing the image. One method resized the full scan into a standard format. Another divided each slice into smaller tiles. A third kept the image at its original level of detail and extracted small regions directly from that native scan. We also evaluated a step called contrast neutralization, which reduces brightness differences between scan types, so the model focuses more on anatomy and less on visual appearance.
Native resolution features excelled for contrast retrieval (97% accuracy), even with a 10% search database. For orientation detection, contrast neutralization with full-FOV features provided the best performance (78% accuracy).2
Our findings suggest that a large reusable model is not necessarily reliable solely because it has been trained on large datasets. In MRI, performance may depend on aligning image representation with the specific task. Fine local detail may be helpful when the task depends on subtle signal differences, while reducing appearance variation may be helpful when the task depends more on overall anatomy and viewing direction.
Scaling quality checks without adding review burden
A persistent challenge in clinical imaging is that quality checking often depends on expert review. While effective at small scale, this approach becomes more difficult when automated tools generate large numbers of outputs. This raises the question of whether there is a practical way to check results without overloading users.
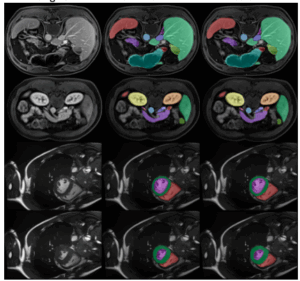
We explored this problem in brain MRI slice placement. Slice placement involves selecting the correct imaging plane so that a structure of interest is captured appropriately. In this study, we focused on landmarks such as the optic nerve, ACPC, hippocampus, and IAC, which require careful positioning for clinical use.
The method used two stages. First, a one-shot segmentation model localized the anatomy. Segmentation refers to identifying the part of the image the system should focus on. Once that region was identified, a vision foundation model (DINOv2) compared the cropped image against a set of expert-approved template images. The output was a similarity score representing how closely the new image matched a reference example.
This automated quality-checking framework showed agreement with expert review under the study conditions. Overall agreement reached 88.2 percent, and the method demonstrated the ability to distinguish between acceptable and unacceptable cases.
The method is not intended to replace expert judgment. It is designed to flag uncertain or failed cases for human oversight. We also observed that pathology cases may require different templates, as diseased anatomy can differ from healthy anatomy. This highlights that a quality-checking system may be more appropriate when it supports routine triaging, while clearly identifying cases that require human review.
Helping Cardiac MRI get the timing right
Late gadolinium enhancement (LGE) is a cardiac MRI technique used after contrast injection to highlight scar or damaged tissue. A key step in this exam is selecting the inversion time (TI), which determines how normal tissue appears relativeto abnormal tissue. If selected appropriately, interpretation may be easier; if not, contrast between tissues may be reduced.
Because TI selection often depends on operator judgment, it is a step that may benefit from additional support. In this work, we developed an image-based method that predicts a candidate TI while also providing an indication of confidence.
The method combines several components. It identifies the relevant region of the image, predicts a timing point, evaluates the suitability of scout images, and includes a confirmation step to assess whether the selected result is consistent.
Across multicenter data, the method helped differentiate valid scout images from inadequate ones and provided feedback indicating whether the suggested result could be accepted, should be reviewed, or whether manual prescription was required due to image limitations.
These results illustrate how automation may support a clinically sensitive step while maintaining transparency. Rather than replacing user judgment, the method is intended to provide guidance along with confidence information.
Aligning Images more reliably across use cases
Registration is the process of aligning two images so that the same anatomy corresponds across scans. This is often needed when comparing images acquired at different times or with different techniques. The task becomes more complex when anatomy changes or when images come from different modalities such as MRI and CT.
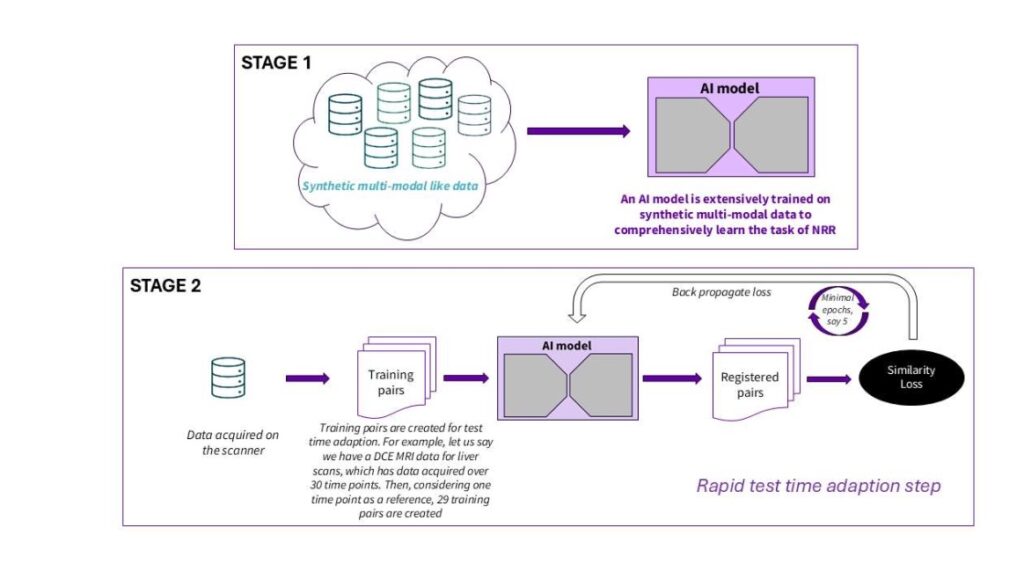
Many registration methods perform well under specific conditions but may be less reliable when those conditions change. We explored a two-stage framework for 3D non-rigid registration, where alignment can deform rather than simply shift.
The first stage involved training on synthetic multimodal image pairs. The second stage applied test-time adaptation, where the model adjusts to the specific image pair being aligned.
We evaluated this approach in two settings: respiratory motion correction in liver dynamic contrast-enhanced MRI, and alignment between pelvic MRI and CT. In liver imaging, the method recovered more physiologically plausible signal behavior under study conditions. In pelvic imaging, it supported alignment between modalities.
These findings suggest a potential direction toward registration approaches that can adapt to varying anatomy and imaging conditions rather than relying only on fixed training scenarios.
Correcting distortion in fast MRI scans
Echo planar imaging (EPI) is widely used due to its speed, particularly in diffusion-weighted imaging. However, it is prone to geometric distortion, where anatomy may appear shifted or warped, especially in areas with magnetic field inhomogeneity.
In clinical contexts, distortion can affect interpretation and comparison across scans. A corrected image is most useful when anatomical structure is represented accurately.
We explored a hybrid method combining model-based reconstruction with learned priors. Model-based reconstruction maintains consistency with MRI signal physics, while learned priors provide data-driven guidance on plausible image structure.
The system estimates magnetic field variation and applies a correction that remains consistent with the acquired signal. We evaluated the method across phantom, prostate, and brain diffusion imaging. In prostate imaging, distortion was reduced in regions commonly affected. In brain imaging, distortion was reduced in areas near the sinuses, with more anatomically plausible structures observed.
This work highlights the importance of balancing visual appearance with anatomical accuracy. The method is intended to improve both aspects by combining physics-based and data-driven approaches.
A broader move toward dependable MRI workflows
Taken together, these studies suggest a consistent direction. Rather than focusing only on performance in controlled benchmarks, the work explores how MRI methods behave under real-world variability and how they may be made more dependable.
In one case, this involved adapting image preparation to the task. In another, it involved supporting quality review while maintaining human oversight. In cardiac imaging, it involved assisting parameter selection with confidence estimates. In registration, it involved adapting to varying anatomy and modalities. In distortion correction, it involved improving geometry while maintaining physical consistency.
A common theme is that an MRI method may provide greater clinical utility when it performs consistently under variation, not only under ideal conditions.
- Concept only. This work is in a concept phase and represents ongoing research. It is not a product, may never become a product, and is not for sale. Any results described are preliminary, subject to change, and have not been cleared or approved by the U.S. FDA or any other global regulator for commercial availability. ↩︎
- https://arxiv.org/pdf/2304.07193 ↩︎