Imagine a future where breast cancer screening tools may potentially be developed on top of a single, mammography-trained vision backbone, reducing the need to start from scratch for every new computer-aided diagnosis feature. That is the objective behind our latest research: MammoDINO1, a vision foundation model tailored specifically for mammography and pre-trained with a novel self-supervised learning framework.
The idea is simple: if the base model learns strong, mammography-specific visual representations from large-scale clinical data without manual labels, developers may be able to adapt it to multiple downstream tasks like cancer detection, lesion localization, BI-RADS prediction, breast density classification, and lesion type prediction, with potentially less dependence on expensive annotation.
The challenge
Breast cancer is among the most commonly diagnosed cancers among women in the United States, and mammography remains the primary screening tool. Yet interpreting mammograms is difficult for several reasons:
• High inter-reader variability, where different radiologists may interpret subtle findings differently
• Early-stage lesions that can be faint and visually ambiguous
• Breast density that can obscure abnormalities and increase false negatives
While CAD tools can support radiologists, many existing systems rely on supervised learning, which depends on large, carefully labeled datasets. In mammography, high-quality labels are costly and time-consuming to produce at scale.
Self-supervised learning (SSL), where models learn from unlabeled data by solving proxy objectives, is a promising research approach. However, generic SSL pipelines often underperform in mammography:
• Random cropping can oversample non-informative background regions and under-sample clinically meaningful breast tissue.
• Standard 2D contrastive objectives can miss cross-slice continuity in tomosynthesis (DBT) volumes / quasi-3D DBT (i.e. we pre-train on 2D slices from DBT volumes), where anatomy changes smoothly across adjacent slices.
A mammography-focused foundation model
To address these gaps, we pre-trained MammoDINO on a large mammography-only dataset of more than 1.4 million 2D images2 from clinical sites across the United States and Europe. This focus on mammography is intentional: rather than learning broadly across many modalities, the model is designed to align with the anatomical and acquisition characteristics that are specific to screening mammograms and DBT.
The training corpus includes:
• 42,000 reconstructed 2D mammograms, including standard mediolateral oblique and craniocaudal views
• 1.35 million 2D slices derived from DBT volumes
To reduce variation across acquisition sites while preserving clinically relevant contrast, we standardize inputs with a lightweight preprocessing pipeline, including intensity normalization and contrast-limited adaptive histogram equalization (CLAHE).
Methodology
MammoDINO is built on a Vision Transformer (ViT) encoder and trained with a DINOv2-based student-teacher SSL framework [1]. In student-teacher SSL, a teacher network produces target representations that the student network learns to match, enabling learning without manual labels. The outcome is a visual encoder that can be transferred to downstream research tasks with lightweight fine-tuning.
Two mammography-aware design choices are intended to better align the SSL training signal with clinical imaging:
Breast tissue-aware augmentation
Standard random crops can waste training signal on background. We use breast tissue-aware data augmentation so that the augmented crops and masked regions used for training are constrained to clinically meaningful breast tissue areas.
In practice, we estimate breast tissue regions directly from the image and enforce a minimum breast-tissue fraction in sampled crops. We apply the same idea to masking at the patch level, prioritizing masking over breast tissue so the model is consistently trained on clinically meaningful structure.
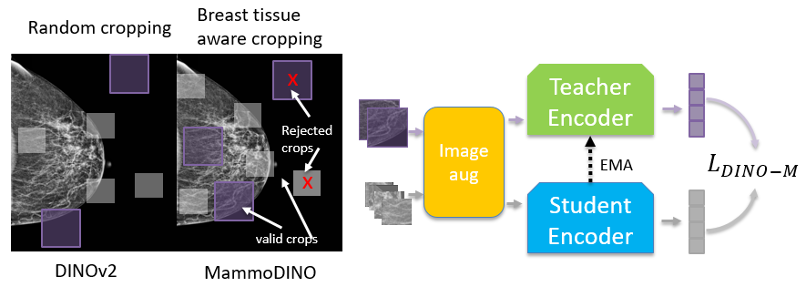
Figure 1(a) illustrates breast tissue-aware crops.
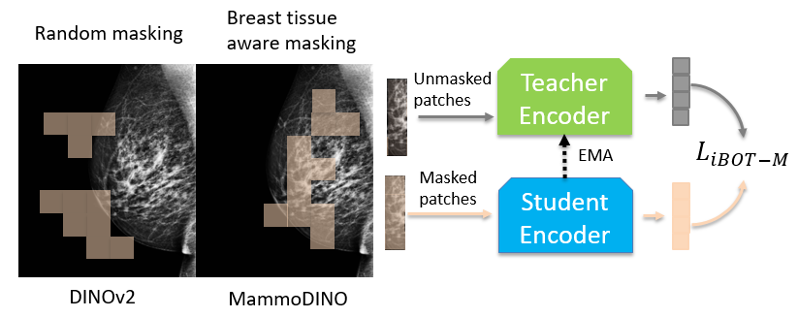
Figure 1(b) illustrates breast tissue-aware masking.
DBT adjacent slice loss
DBT volumes consist of closely spaced 2D slices where anatomy changes gradually along the depth axis. To reflect this structural coherence, we introduce a DBT adjacent slice loss that aligns model predictions for nearby slices from the same DBT volume.
Concretely, we sample nearby slices from the same DBT volume, apply conservative tissue-aware transforms, and train the student representation for one slice to align with the teacher representation of its neighbor. This encourages features that remain stable across depth, reflecting the smooth anatomical transitions in DBT.
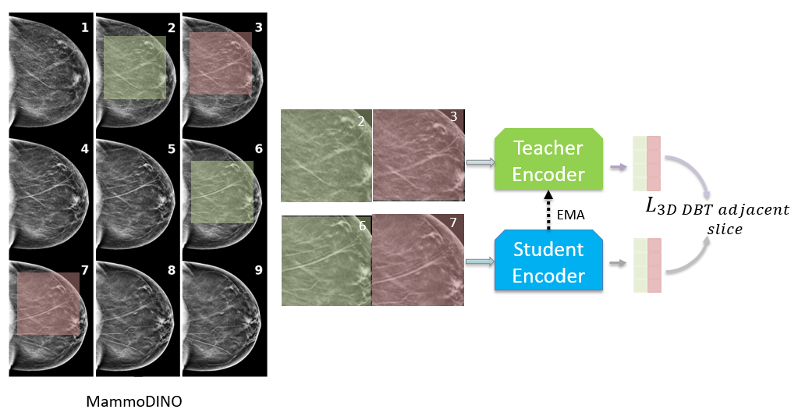
Figure 1(c) shows the DBT adjacent slice loss concept.
Together, these choices are designed to encourage MammoDINO to learn more discriminative and clinically meaningful representations, using training signals that reflect both the physiology and geometry of mammography.
Benchmark performance
We evaluated MammoDINO across five major public benchmark datasets: RSNA, VinDr-Mammo, DDSM, CMMD, and CDD-CESM. The model was tested on downstream research tasks commonly studied in screening and diagnostic workflows:
• Breast cancer detection
• Lesion detection
• BI-RADS score prediction
• Breast density prediction
Baselines included supervised CNNs (ResNet50 [2], ConvNeXt [3]), generic self-supervised vision encoders (DINOv2), and medical domain models with varying supervision strategies (RadDINO [4], BiomedCLIP [5], MammoCLIP [6]). For a consistent comparison, we adapt each backbone with the same lightweight classification head and fine-tune per dataset and task.
Breast cancer and lesion detection
On VinDr-Mammo, MammoDINO achieved an AUC of 0.918 for cancer detection under the reported experimental conditions, compared with ResNet50 (0.554) and MammoCLIP (0.809). AUC, or Area Under the ROC Curve, measures how well a model separates positive from negative cases across thresholds.
On DDSM, it achieved 0.932 AUC for lesion detection under the reported study setup, compared with ResNet50 (0.919) and the baseline DINOv2 encoder (0.902).
BI-RADS score prediction
BI-RADS is a standardized clinical reporting system used in breast imaging. For BI-RADS prediction on CMMD, MammoDINO reached an F1 score of 0.762 under the study protocol, compared with DINOv2 (0.704) and ResNet50 (0.633). F1 balances precision and recall, making it useful when class distributions are imbalanced.
Breast density classification
Breast density is clinically important because dense tissue can mask early lesions. On RSNA for multi-class density prediction, MammoDINO scored 0.759 under the reported evaluation framework, compared with MammoCLIP (0.748) and RadDINO (0.740).
Figure 2 provides a radar plot summarizing performance across models and tasks.
One practical takeaway from the broader benchmark suite is that gains are not perfectly uniform across every dataset-task pair, but the overall trend in the reported experiments indicates that mammography-aware pre-training is associated with strong transfer performance across tasks and benchmarks.
Implications and research context
This work suggests that modality-specific, anatomically aware SSL may support the development of stronger research foundations for breast imaging CAD exploration. By aligning pre-training objectives with mammography’s acquisition characteristics, MammoDINO represents a reusable visual backbone for multiple research tasks.
We also observe in the reported ablation analyses that the mammography-aware design choices contribute to performance differences. When components are evaluated separately, both tissue-aware sampling and the DBT adjacent slice objective are associated with performance improvements across datasets on tasks such as cancer detection, lesion detection, and BI-RADS prediction. Qualitatively, feature visualizations suggest that the learned representations concentrate on clinically meaningful tissue regions and highlight areas of interest, consistent with the motivation behind tissue-aware training.
Even though MammoDINO is instantiated for mammography, the underlying training schema is intended as a research framework. The same concept — using anatomy-aware augmentation and structure-aware consistency losses — could potentially be explored for other imaging modalities, including CT and MRI, where the geometry of the data and the clinical meaning of regions-of-interest may shape the learning signal.
Conclusion
MammoDINO advances mammography-focused foundation modeling research by combining large-scale, mammography-only pre-training with SSL objectives designed around breast tissue relevance and DBT slice continuity. The result is a scalable encoder that demonstrated strong performance across public benchmarks and research tasks under the reported study conditions, and represents a potential pathway for continued CAD research in breast cancer screening.
References
[1] Oquab, Maxime, et al. “DINOv2: Learning robust visual features without supervision.” arXiv:2304.07193 (2023).
[2] He, Kaiming, et al. “Deep residual learning for image recognition.” CVPR (2016).
[3] Woo, Sanghyun, et al. “ConvNeXt V2: Co-designing and scaling convnets with masked autoencoders.” CVPR (2023).
[4] Perez-Garcia, Fernando, et al. “Exploring scalable medical image encoders beyond text supervision.” Nature Machine Intelligence 7.1 (2025): 119-130.
[5] Zhang, Sheng, et al. “BiomedCLIP.” arXiv:2303.00915 (2023).
[6] Chen, Xuxin, et al. “Mammo-CLIP.” arXiv:2404.15946 (2024).
[7] Reza, A. M. “Realization of the contrast limited adaptive histogram equalization (CLAHE) for real-time image enhancement.” Journal of VLSI Signal Processing Systems (2004).
- Concept only. May never become a product. Not for Sale. Not cleared or approved by the U.S. FDA or any other global regulator for commercial availability. ↩︎
- Source: Unveiling MammoDINO: an anatomically aware vision foundation model for mammography (arXiv: https://arxiv.org/pdf/2510.11883) ↩︎